Sponsor:
PnetCDF development was sponsored by the Scientific Data Management Center (SDM) under the DOE program of Scientific Discovery through Advanced Computing (SciDAC). It was also supported in part by National Science Foundation under the SDCI HPC program award numbers OCI-0724599 and HECURA program award numbers CCF-0938000. PnetCDF development is currently supported by the Scientific Data, Analysis, and Visualization (SDAV) Institute under the DOE SciDAC program (Award Number DE-SC0007456).Project Team Members:
Northwestern University
- Wei-keng Liao
- Alok Choudhary
- Kai-Yuan Hou (Ph.D. student graduated in 2022)
- Seung Woo Son (formerly a postdoc, now an Assistant Professor at UMass Lowell)
- Kui Gao (formerly postdoc, now Dassault Systèmes Simulia Corp.
- Jianwei Li (since graduated, now Bloomberg L.P.)
Argonne National Laboratory
- Rob Latham
- Rob Ross
- Rajeev Thakur
- William Gropp(now UIUC)
PnetCDF

News:
- Release of PnetCDF version 1.14.0 is available on November 11, 2024.
- We have been working with the netCDF team at Unidata to integrate CDF-5 and PnetCDF features into netCDF 4.4.0. See release note of 4.4.0 for more information.
- PnetCDF was used in a large-scale simulation of hurricane Sandy running on the Blue Waters supercomputer at NCSA in 2013.
Contents:
- What is netCDF?
- Design strategy for parallelizing netCDF
- New features added in PnetCDF
- Interoperability with NetCDF-4
- Download source codes
- User documents and example programs
- I/O benchmarking programs
- Publications
- User community
- Contact
What is netCDF?
NetCDF (Network Common Data Form) is an I/O library that supports the creation, access, and sharing of scientific data.- File formats -- Self-describable, machine-independent file formats (CDF and HDF5) are used to store multi-dimensional array-oriented data together with its attributes (such as annotations.) The data layout in CDF files follows the canonical order of the arrays.
- Application Programming Interfaces (APIs) -- Set of Fortran, C, C++, and Java functions are available in the netCDF software releases for accessing the data stored in the files (in CDF and HDF5 formats). The APIs are used to define dimensions, variables, attributes of variables, and perform data read/write to the files.
Starting from versions 4.0 and 4.1, Unidata's netCDF supports parallel I/O through HDF5 and PnetCDF underneath, respectively. Through PnetCDF, netCDF-4 programs can access files in the classical CDF formats in parallel. Similarly, through HDF5, netCDF-4 programs can access files in HDF5 format.
Design Strategy for Parallelizing NetCDF:
The goal of PnetCDF is to provide high-performance parallel I/O to the applications by enabling all client processes to access a shared file in parallel. To ensure the performance and portability, PnetCDF is built on top of MPI-IO. The PnetCDF APIs incorporate the parallel semantics following the MPI (Message Passing Interfaces) standard and provide backward compatibility with the classical netCDF file formats: CDF (or CDF-1), CDF-2, and CDF-5. Figure 1 compares the data access methods between the sequential netCDF and PnetCDF.
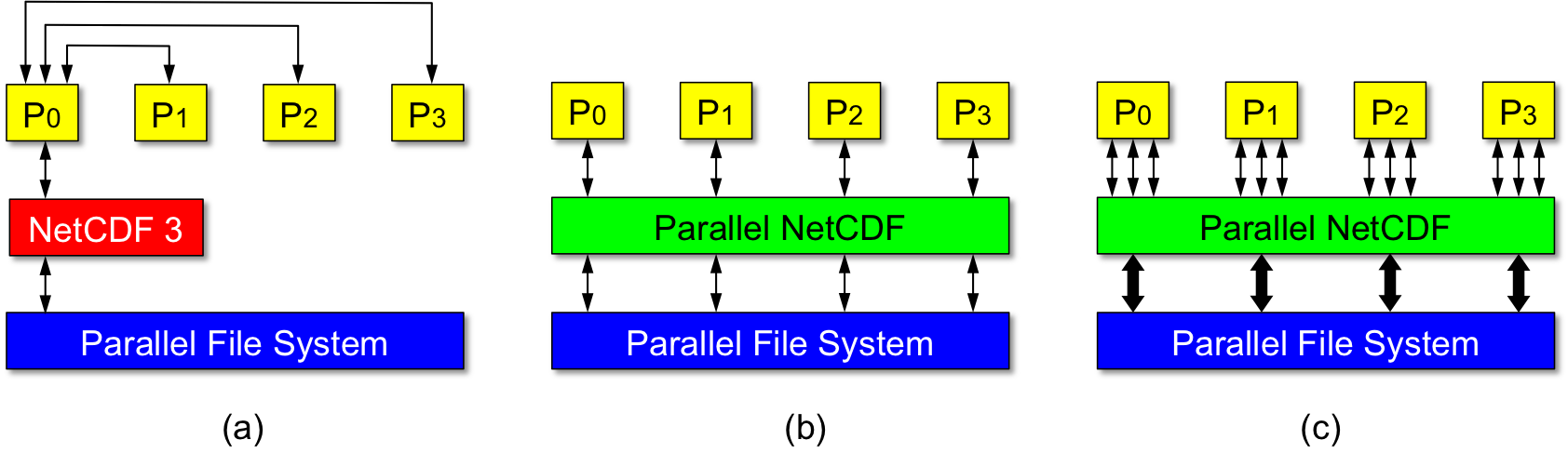
Below describes the design principle of PnetCDF.
- Minimize the changes to the netCDF API syntax --
In order for easy code migration from sequential netCDF to PnetCDF,
PnetCDF APIs mimic the syntax of the netCDF APIs with only minor
changes to add parallel I/O concept. These changes are highlighted as
follows.
- All parallel APIs are named after the original netCDF APIs with
"ncmpi_" prefix for C/C++, "nfmpi_" for Fortran 77, and
"nf90mpi_" for Fortran 90. For example,
int ncmpi_put_vara_float(int ncid, /* dataset ID */ int varid, /* variable ID */ const MPI_Offset start[], /* [ndims] */ const MPI_Offset count[], /* [ndims] */ float *buf) /* user buffer */ - An MPI communicator and an MPI_Info object are added to the
argument list of the open/create APIs. The communicator defines the
set of processes that will access the netCDF file in parallel. The
info object allows users to provide I/O hints for PnetCDF and
MPI-IO to further improve performance (e.g. file alignment for file
header size and starting offsets for variables, and the MPI-IO
hints.) An example is
int ncmpi_open(MPI_Comm comm, /* the group of MPI processes sharing the file */ const char *path, int omode, MPI_Info info, /* PnetCDF and MPI-IO hints */ int *ncidp) - PnetCDF defined two data modes, collective and independent,
which correspond to MPI collective and independent I/O operations.
Similar to MPI naming convention, all collective APIs carry an
extra suffix "_all". The independent I/O mode is wrapped by the
calls of ncmpi_begin_indep_data() and ncmpi_end_indep_data().
While in collective data mode, only calls to collective APIs are
allowed. Similarly, only independent APIs are allowed in
independent mode. The API ncmpi_put_vara_float() shown above is an
independent API and the collective counterpart is:
int ncmpi_put_var_float_all(int ncid, /* dataset ID */ int varid, /* variable ID */ const MPI_Offset start[], /* [ndims] */ const MPI_Offset count[], /* [ndims] */ float *buf) /* user buffer */ - For the API arguments related to variable sizes or their
accesses that are of type size_t, PnetCDF replaces it with
MPI_Offset. For example, the arguments start[] and count[] in
the above APIs are of MPI_Offset data type vs. size_t in
netCDF. Another example is the API defining a dimension,
given below. The arguments of type ptrdiff_t are also changed
to MPI_Offset, including arguments stride[] and imap[] in vars
and varm API families.
int ncmpi_def_dim(int ncid, /* dataset ID */ const char *name, /* dimension name string */ const MPI_Offset len, /* length of dimension */ int *dimidp) /* returned dimension ID */
- All parallel APIs are named after the original netCDF APIs with
"ncmpi_" prefix for C/C++, "nfmpi_" for Fortran 77, and
"nf90mpi_" for Fortran 90. For example,
- Support large files -- PnetCDF supports CDF-2 file format (by setting NC_64BIT_OFFSET flag when creating a new file). With CDF-2 format, even on 32-bit platforms one can create netCDF files of size greater than 2GB.
New features added in PnetCDF:
- Support large variables -- PnetCDF supports CDF-5 file format. With CDF-5 format, large sized array variables with more than 4 billion elements can be created in a file.
- Support additional data types -- New data types introduced in CDF-5 are NC_UBYTE, NC_USHORT, NC_UINT, NC_INT64, and NC_UINT64.
- PnetCDF I/O hints -- PnetCDF I/O hints, nc_header_align_size and nc_var_align_size, allows users to set a customized file header size and starting file offsets of non-record variables. File layout alignment has been known to cause significant performance impact on parallel file system. The hint nc_header_align_size can be used to reserve a sufficiently large space for file header, in case more metadata is to be added to an existing netCDF file. The common practice for setting the hint nc_var_align_size is the file system striping size. An example program can be found in hints.c.
- Flexible APIs for non-contiguous user buffers --
In addition to the support of existing netCDF functionality, another new
set of APIs, called flexible APIs, are available to make use of MPI
derived datatypes to describe a complex memory layout for user I/O buffers.
On the contrary, traditional netCDF APIs allow only contiguous data buffers.
Example programs can be found in
flexible_api.c and
flexible_api.f.
An example API is given below.
int ncmpi_put_vara_all(int ncid, int varid, const MPI_Offset start[], /* [ndims] */ const MPI_Offset count[], /* [ndims] */ void *buf, /* user I/O buffer */ MPI_Datatype bufcount, /* number of buftype elements in buf */ MPI_Datatype buftype) /* MPI derived data type */ - vard APIs -- takes an argument of MPI derived data type that
describes the file access layout, as opposed to vara and vars APIs that use
start[] and count[]. Through this API family, users can access complex
non-contiguous file space, no longer limited to a subarray layout.A An
example API is
int ncmpi_put_vard_all(int ncid, /* dataset ID */ int varid, /* variable ID */ const MPI_Datatype filetype, /* file access layout */ const void *buf, const MPI_Offset bufcount, const MPI_Datatype buftype); /* buffer layout in memory */ - varn APIs -- for making multiple requests to the same variable
Conventional netCDF APIs (i.e. var, var1, vara, vars, and varm) allow one
request to a variable per API call. A new set of APIs, named varn, is
introduced to allow making multiple requests with arbitrary locations to a
single variable. See the example program
put_varn_float.c).
An example API is
int ncmpi_put_varn_float_all(int ncid, /* dataset ID */ int varid, /* variable ID */ int num, /* number of requests */ MPI_Offset* const *start, /* [num][ndims] list of start offsets */ MPI_Offset* const *counts, /* [num][ndims] list of access counts */ float *buf); /* buffer pointer */ - mput/mget APIs -- for making multiple requests to different
variables. This API family lets a single API call to complete multiple
subarray requests to the same and/or different variables. See the example
program mput.c).
An example API is
int ncmpi_mput_vara_all(int ncid, /* dataset ID */ int nvars, /* number of variables */ int varids[], /* [nvars] list of variable IDs */ MPI_Offset* const *starts, /* [nvars][ndims] list of start offsets */ MPI_Offset* const *counts, /* [nvars][ndims] list of access counts */ void *bufs[], /* [nvars] list of buffer pointers */ const MPI_Offset bufcounts[], /* [nvars] list of buffer counts */ const MPI_Datatype buftypes[]); /* [nvars] MPI derived datatypes describing bufs */ - iput/iget APIs -- for request aggregation.
Nonblocking APIs (ncmpi_iput_xxx/ncmpi_iget_xxx) are designed to aggregate
smaller I/O requests into large ones for better performance. A common
practice is to first post multiple nonblocking calls and then use a single
call to ncmpi_wait_all() to complete the I/O transaction. Aggregation
applies to requests to the same variables as well as across different
variables. See examples/README for example programs.
int ncmpi_iget_vara_int64(int ncid, int varid, const MPI_Offset start[], /* [ndims] */ const MPI_Offset count[], /* [ndims] */ void *buf, /* user I/O buffer */ int *request_id); - bput APIs -- for buffered write.
Buffered write APIs (ncmpi_bput_xxx) is another set of nonblocking APIs
that caches the request data in an internal buffer, so that the user buffer
can be reused of freed once the posting call returns. This API set is in
contract to the iput/iget API family that requires user buffer untouched
before the wait API is completed. User programs must first call
ncmpi_buffer_attach() to specify an amount of internal buffer that can be
used by PnetCDF to aggregate the write requests. Example programs can be
found in examples/tutorial/pnetcdf-write-buffered.c and examples/tutorial/pnetcdf-write-bufferedf.f90.
int ncmpi_bput_vara_int64(int ncid, int varid, const MPI_Offset start[], /* [ndims] */ const MPI_Offset count[], /* [ndims] */ void *buf, /* user I/O buffer */ int *request_id);
Interoperability with NetCDF-4
- Starting from version 4.1, netCDF-4 program can perform parallel I/O
on the classic CDF-1 and CDF-2 files through PnetCDF. This is done by
passing file create flag NC_PNETCDF to nc_create_par(), for instance,
nc_create_par(filename, NC_PNETCDF|NC_CLOBBER, MPI_COMM_WORLD, info, &ncid);
Flag NC_PNETCDF has been deprecated since NetCDF 4.6.3 and using it is no longer required. For example,
nc_create_par(filename, NC_CLOBBER, MPI_COMM_WORLD, info, &ncid); - Note some new APIs of PnetCDF are not available in netCDF-4 yet. These APIs are "flexible", "nonblocking", "mput/mget", "vard", and "varn".
- Example programs in C and Fortran are available below that show how to
enable parallel I/O using netCDF-4 APIs. They can be easily changed to
use either PnetCDF or HDF5 libraries underneath. At the top of each
example program, the instructions for configuring netCDF to enable
PnetCDF option are provided, as well as for compiling and running the
example.
- nc4_pnc_put.c is an example program that makes a call to nc_put_vara_int() to write subarrays to a 2D integer array in parallel. In this example, the data partition pattern among processes is in a block fashion along both X and Y dimensions.
- nc4_pnc_get.c is a read example program, the counterpart of nc4_pnc_put.c.
- Fortran version of the above examples can be found in nc4_pnc_put_vara.f and nc4_pnc_get_vara.f
- coll_perf_nc4.c is an I/O performance benchmark program that reports the aggregate bandwidth of writing 20 three-dimensional arrays of integer type in parallel. The data partitioning pattern used is in block-block-block along the three dimensions. This program also reports the performance of using HDF5+MPI-IO method.
Download Source Codes
- The latest stable version 1.14.0 was released on November 11, 2024.
- Please visit Download page for downloading current and prior releases.
- Instructions for building PnetCDF library are provided in the file INSTALL come with the source code release. PnetCDF uses Autoconf tools which is usually smart enough to detect the required software by just running command "./configure". However, if you encounter problems, there are also several build recipes for specific platforms: README.IBM, README.CRAY, and README.SGI.
- Source code repository -- Users are recommended to use the release versions for production runs but also welcomed to try the latest features under development. https://github.com/Parallel-NetCDF/PnetCDF
User Documents and Example Programs
- PnetCDF C Interface Guide is based on the netCDF C interface Guide.
- PnetCDF Q&A contains a few tips for achieving better I/O performance.
- A tutorial
with use cases of popular parallel I/O strategies:
- I/O from the master process
- one file per process
- parallel I/O on a shared file
- using non-contiguous I/O buffer
- using non-blocking I/O, and
- using buffered APIs
- Other example programs
- Read/write variables in a matrix-transposed fashion: transpose.c, transpose.f, transpose.f90, transpose.cpp
- Using fill mode APIs in PnetCDF (note the difference from netCDF for record variables): fill_mode.c, fill_mode.f, fill_mode.f90, fill_mode.cpp
- Using I/O buffers in memory that have ghost cells: ghost_cell.c, flexible_api.c, flexible_api.f, flexible_api.f90, flexible_api.cpp
- Read/write when the data partitioning pattern is a block-cyclic fashion: block_cyclic.c, block_cyclic.f, block_cyclic.f90, block_cyclic.cpp
- Uisng varn APIs to read/write multiple sub-arrays in a single API call: put_varn_float.c, put_varn_int.c, put_varn_float.f, put_varn_int.f, put_varn_float.f90, put_varn_int.f90, put_varn_float.cpp put_varn_int.cpp
- Nonblocking version of varn API: bput_varn_uint.c, i_varn_int64.c, bput_varn_int8.f, i_varn_real.f,
- See README and the beginning of each example program for additional descriptions
- All tutorial and example programs are available in the PnetCDF releases, under the directory named "examples".
Under Development
PnetCDF is constantly updated with new features and performance improvement methods to meet the high-performance computing demands. The following list some task currently under development.- Subfiling -- a scheme that can divide a large multi-dimensional global array into smaller subarrays, each saved in a separate netCDF file, named subfile. The subfiling scheme can decrease the number of processes sharing a file, so to reduce the file access contention, an overhead the file system pays to maintain data consistency. Subfiling is available in PnetCDF release 1.4.1.
I/O Performance Benchmarking Programs
- PnetCDF in BTIO
- BTIO is the I/O part of NASA's NAS Parallel Benchmarks (NPB) suite.
- btio-pnetcdf-1.1.1.tar.gz (SHA1 checksum: 7a1380e496f7c328d7605c560c94bbbd84409cfc)
- PnetCDF in S3D-IO
- S3D is a continuum scale first principles direct numerical simulation code developed at Sandia National Laboratory. S3D-IO is its I/O kernel.
- s3d-io-pnetcdf-1.2.1.tar.gz (SHA1 checksum: c2a5fc2175d9836ce39f76c8c8bbcd9065782d1d)
- PnetCDF in GCRM-IO
- Global Cloud Resolving Model (GCRM) developed at Colorado State University, is a climate application framework designed to simulate the circulations associated with large convective clouds.
- The I/O module in GCRM uses Geodesic I/O library (GIO) developed at Pacific Northwest National Laboratory.
- Note that the GCRM I/O kernel benchmark program written in Fortran is included in the GIO source release. The tar ball downloadable here contains the C version converted from the GIO's Fortran version.
- gcrm-io-pnetcdf-1.0.0.tar.gz (SHA1 checksum: 32bd510faf4fcef3edeb564d3885edac21f8122d)
- PnetCDF in FLASH-IO
- FLASH is a block-structured adaptive mesh hydrodynamics code developed mainly for the study of nuclear flashes on neutron stars and white dwarfs.
- The PnetCDF method is developed based on the FLASH I/O benchmark suite and is included in the PnetCDF release starting from v1.4.0.
Publications
- Seung Woo Son, Saba Sehrish, Wei-keng Liao, Ron Oldfield, and Alok Choudhary. Dynamic File Striping and Data Layout Transformation on Parallel System with Fluctuating I/O Workload. In the Workshop on Interfaces and Architectures for Scientific Data Storage, September 2013.
- Rob Latham, Chris Daley, Wei-keng Liao, Kui Gao, Rob Ross, Anshu Dubey, and Alok Choudhary. A Case Study for Scientific I/O: Improving the FLASH Astrophysics Code. Computer and Scientific Discovery, 5, March 2012.
- Kui Gao, Chen Jin, Alok Choudhary, and Wei-keng Liao. Supporting Computational Data Model Representation with High-performance I/O in Parallel netCDF. In the IEEE International Conference on High Performance Computing, December 2011.
- Kui Gao, Wei-keng Liao, Arifa Nisar, Alok Choudhary, Robert Ross, and Robert Latham. Using Subfiling to Improve Programming Flexibility and Performance of Parallel Shared-file I/O. In the Proceedings of the International Conference on Parallel Processing, Vienna, Austria, September 2009.
- Kui Gao, Wei-keng Liao, Alok Choudhary, Robert Ross, and Robert Latham. Combining I/O Operations for Multiple Array Variables in Parallel netCDF. In the Proceedings of the Workshop on Interfaces and Architectures for Scientific Data Storage, held in conjunction with the IEEE Cluster Conference, New Orleans, Louisiana, September 2009.
- Jianwei Li, Wei-keng Liao, Alok Choudhary, Robert Ross, Rajeev Thakur, William Gropp, Rob Latham, Andrew Siegel, Brad Gallagher, and Michael Zingale. Parallel netCDF: A Scientific High-Performance I/O Interface. In the Proceedings of Supercomputing Conference, November, 2003.
User Community:
- Community Multiscale Air Quality Model
(CMAQ)
An active open-source development project of the U.S. EPA Atmospheric Science Modeling Division that consists of a suite of programs for conducting air quality model simulations.
Organization: CMAS Center
Reference: What's New with the I/O API
- Community Earth System Model
(CESM)
A global atmosphere model for use by the wider climate research community.
Organization: UCAR
People: Jim Edwards
The program uses PIO library for parallel I/O operations, which includes PnetCDF method. CESM software prerequisites - RAMSES-GPU
A general-purpose Hydrodynamics (HD) and Magneto-hydrodynamics (MHD) simulation code primarily written for astrophysics applications.
Organization: Maison de la Simulation and CEA/Sap
People: Pierre Kestener and Sebastien Fromang
Reference: Weak scaling measured on OLCF/TITAN - Data Services for the Global Cloud Resolving Model (GCRM)
A global atmospheric circulation model with a grid-cell spacing of approximately 3 km, capable of simulating the circulations associated with large convective clouds.
using PnetCDF, Fortran 90, C++
Organization: Pacific Northwest National Laboratory
People: Karen Schuchardt
Reference: B. Palmer, A. Koontz, K. Schuchardt, R. Heikes, and D. Randall. Efficient Data IO for a Parallel Global Cloud Resolving Model Environ. Model. Softw. 26, 12 (December 2011), 1725-1735. DOI=10.1016/j.envsoft.2011.08.007. - NCAR Community Atmosphere Model (CAM)
using Parallel netCDF, ZioLib
Platforms: IBM SP3, SP4, SP5, BlueGene/L, Cray X1E
File systems: GPFS, PVFS2, NFS
Organization: Department of Atmospheric Sciences, National Taiwan University
People: Yu-heng Tseng (yhtseng at as.ntu.edu.tw)
Reference: Tseng, Y. H. and Ding, C.H.Q. Efficient Parallel I/O in Community Atmosphere Model (CAM), International Journal of High Performance Computing Applications, 22, 206-218, 2008. - Astrophysical Thermonuclear Flashes (FLASH)
using PnetCDF, HDF5, C, User Guide
Platforms: IBM SP, Linux Clusters
Organization: ASCI Flash Center, University of Chicago
People: Brad Gallagher, Katie Antypas
Reference: R. Latham, C. Daley, W. Liao, K. Gao, R. Ross, A. Dubey and A. Choudhary. A Case Study for Scientific I/O: Improving the FLASH Astrophysics Code. In the Computational Science and Discovery, vol. 5, 2012. DOI. 10.1088/1749-4699/5/1/015001 - ASPECT, parallel VTK
using PnetCDF, C
Platforms: Linux Clusters, Cray X
Organization: ORNL
People: Nagiza Samatova - Atmospheric Chemical Transport Model (ACTM)
using PnetCDF, FORTRAN
Organization: Center for Applied Scientific Computing, LLNL
People: John R. Tannahill - PRogram for Integrated Earth System Modeling (PRISM) Support Initiative
supports netCDF and PnetCDF within the IO library as part of the OASIS4 coupler
using netCDF, PnetCDF (FORTRAN APIs)
Platforms: NEC SX, Linux Cluster, SGI and others
Organization: C&C Research Laboratories, NEC Europe Ltd.
Contacts for pnetcdf in PRISM: Reiner Vogelsang and Rene Redler
Contacts for pnetcdf users on NEC SX: Joachim Worringen and Rene Redler - Weather Research and Forecast
(WRF) modeling system software
using PnetCDF, FORTRAN, see WRF Installation Best Practices
Organization: National Center for Atmospheric Research (NCAR)
People: John Michalakes
News item: Hurricane Force Supercomputing: Petascale Simulations of Sandy
HWRF (WRF for Hurricanes). - Stony Brook Parallel Ocean Model (sbPOM)
using PnetCDF, FORTRAN
sbPOM is a parallel, free-surface, sigma-coordinate, primitive equations ocean modeling code based on the Princeton Ocean Model (POM) People: Antoni Jordi (toniimedea.uib-csic.es) and Dong-Ping Wang (dpwangnotes.cc.sunysb.edu) - WRF-ROMS (Regional Ocean Model System) I/O Module
using PnetCDF, FORTRAN
Organization: Scientific Data Technologies Group, NCSA
People: MuQun Yang, John Blondin - Portable, Extensible Toolkit for Scientific Computation
(PETSc)
Organization: ANL - The Earth System Modeling Framework (ESMF)
platform: IBM Blue Gene / L
Organization: Scientific Computing Division
National Center for Atmospheric Research
Boulder, CO 80305
People: Nancy Collins, James P. Edwards
User manual for building ESMF with PnetCDF support.
- Parallel Ice Sheet Model (PISM)
The Parallel Ice Sheet Model is an open source, parallel, high-resolution ice sheet model.
Related Links
- Unidata's netCDF
- Message Passing Interface Standard
- PnetCDF -- the official project web page at github.com.
- PnetCDF project web page maintained at Argonne National Laboratory, including software download, users documents, etc.
- High Performance I/O: Parallel netCDF - an article by Forrest Hoffman appears in Linux Magazine.
- Philippe Wauteleta and Pierre Kestener. Parallel IO Performance and Scalability Study on the PRACE CURIE Supercomputer, white paper at Partnership For Advanced Computing in Europe (PRACE), September, 2012. This report compares the performance of PnetCDF, HDF5, and MPI-IO on CURIE supercomputer with Lustre parallel file system using IOR and RAMSES-GPU.
Contact
- User Discussion Mailing List
- Participate the discussion and receive announcement by subscribing the mailing list.
- Mailing list archive.
- Questions about this page
- Please email to Wei-keng Liao <wkliao at eece dot northwestern dot edu>
Acknowledgements:
We are grateful to the following people who provide valuable
comments/discussions to improve our implementation.
Yu-Heng Tseng (LBNL)
Reiner Vogelsang (Silicon Graphics, Germany),
Jon Rhoades (Information Systems & Technology ENSCO, Inc.),
Kilburn Building (University Of Manchester),
Foucar, James G (Sandia National Lab.),
Drake, Richard R (Sandia National Lab.),
Eileen Corelli (Senior Scientist, ENSCO Inc.),
Roger Ting,
Hao Yu,
Raimondo Giammanco,
John R. Tannahill (Lawrence Livermore National. Lab.),
Tyce Mclarty (Lawrence Livermore National. Lab.),
Peter Schmitt, Mike Dvorak (LCRC team, MCS ANL)




